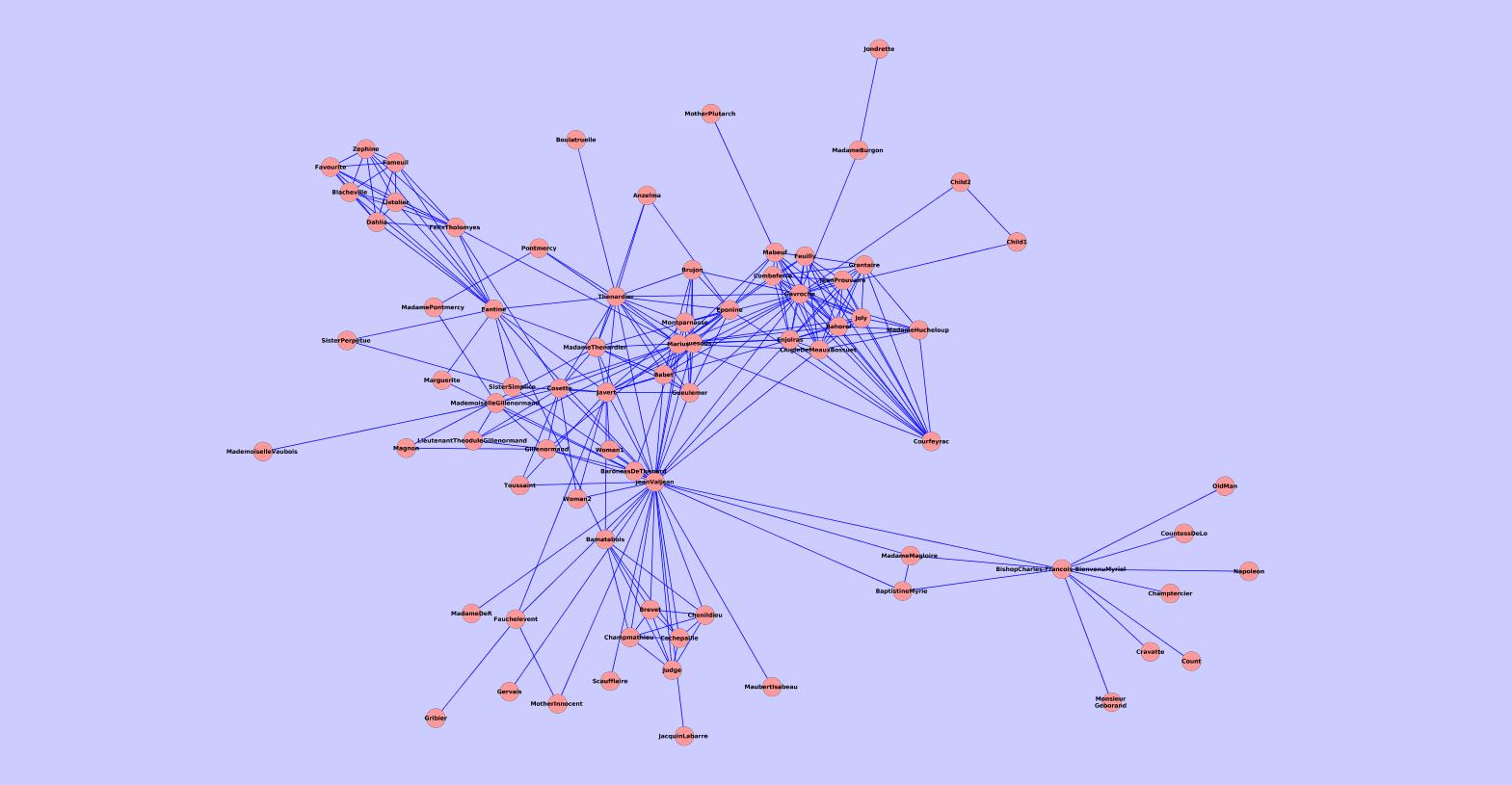
Our method is also fast.ĭense subgraphs and clique structures are not the only interconnectivity pattern with biological significance. In our preliminary analysis on synthetic and real datasets, we find our approach to be competitive with or significantly better than a selection of leading approaches including Metis, MCODE, and spectral clustering. Consequently, almost all methods that propose solutions to this problem are necessarily approximate heuristics. Theoretically, the problem of finding the densest subgraph in a network is known to be NP-hard. Community finding in large networks has become a ubiquitous problem, and many research groups spanning diverse fields have proposed methods. Dense sub-networks of molecules often represent protein complexes and coherent biological processes. 
In molecular interaction networks, groups of densely connected molecules frequently have an important biological interpretation. With estimates on the size of the complete protein interactome for humans and other metazoans topping 650,000 interactions, sophisticated tools are needed to cope with the complexity of biological systems. Taken together these molecular interaction data sets form our picture of the known interactome. Some of these important assays yield protein-protein, protein-DNA, and synthetic lethal genetic interactions. Biochemical assays that annotate associations between pairs of genes and proteins have become increasingly diverse. The vast amount of molecular biology data presents us with new organizational challenges as we seek to extract knowledge from whole-genome experimental assays. Rapid innovation in the field of high-throughput whole-genome biochemistry has revolutionized our understanding of biology. An implementation of our method is freely available for download as a plugin to Cytoscape through our website and through Cytoscape itself. 
We minimize model complexity and generalization error in the Bayesian spirit by integrating out nuisance parameters. Our results suggest that the performance of NeMo is better than or competitive with leading approaches on both real and synthetic datasets. The approach is unique in that we identify both dense network and dense bipartite network structures in a single approach. We present a novel method that combines a unique neighbour-sharing score with hierarchical agglomerative clustering to identify diverse network communities.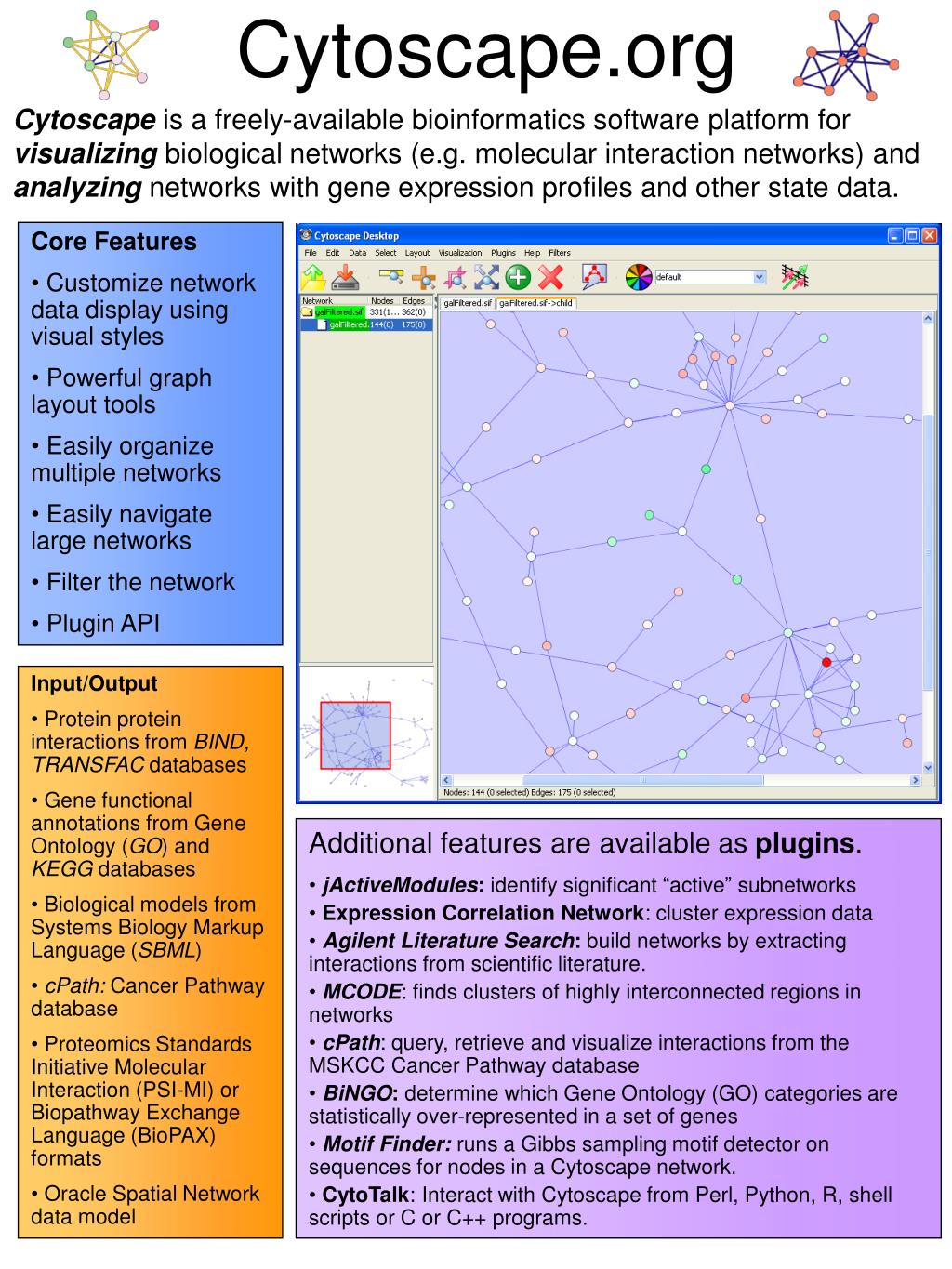 
We apply our method to the CXC chemokine pathway and find a high scoring functional module of 12 disconnected phospholipase isoforms. We test these methods on a collection of synthetically constructed networks and the set of MIPS human complexes. To evaluate the performance of our method (NeMo), we compare it to other widely used tools for community detection including kMetis, MCODE, and spectral clustering. In this work, we present a novel method to identify densely connected and bipartite network modules based on a log odds score for shared neighbours. Previous studies have shown that densely connected network components frequently correspond to community structure and functionally related modules. As the size of the known human interactome grows, biologists increasingly rely on computational tools to identify patterns that represent protein complexes and pathways.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
